Sections in this Chapter:
- Write & run scripts
- Debugging
- Benchmarking
- Vectorization
- Embarrassingly parallelism in R/Python
- Evaluation strategy
- Speed up with C/C++ in R/Python
- A first impression of functional programming
- Miscellaneous
Write & run scripts
In the first Chapter, we are coding within the interactive mode of R/Python. When we are working on the real world projects, using an Integrated development environment (IDE) is a more pragmatic choice. There are not many choices for R IDE, and among them RStudio1 is the best one I have used so far. As for Python, I would recommend either Visual Studio Code2 or PyCharm3. But of course, you could use any text editor to write R/Python scripts.
Let’s write a simple script to print Hello World! in R/Python. I have made a directory chapter2 on my disk, the R script is saved as hello_world.R and the Python script is saved as hello_world.py, inside the directory.
chapter2/hello_world.R
1 print("Hello World!")
chapter2/hello_world.py
1 print("Hello World!")
There are a few ways to run the R script. For example, we can run the script from the console with the r -f filename command. Also, we can open the R interactive session and use the source() function. I would recommend the second approach with source() function. As for the Python script, we can run it from the console.
1 chapter2 $ls
2 hello_world.R hello_world.py
3 chapter2 $r -f hello_world.R
4 > print("Hello World!")
5 [1] "Hello World!"
6
7 chapter2 $r
8 > source('hello_world.R')
9 [1] "Hello World!"1 chapter2 $ls
2 hello_world.R hello_world.py
3
4 chapter2 $python3.7 hello_world.py
5 Hello World!Debugging
Debugging is one of the most important aspects of programming. What is debugging in programming? The programs we write might include errors/bugs and debugging is a step-by-step process to find and remove the errors/bugs in order to get the desired results.
If you are smart enough or the bugs are evident enough then you can debug the program on your mind without using a computer at all. But in general we need some tools/techniques to help us with debugging.
Most of programming languages provide the functionality of printing, which is a natural way of debugging. By trying to place print statements at different positions we may finally catch the bugs. When I use print to debug, it’s feeling like playing the game of minesweeper. In Python, there is a module called logging4 which could be used for debugging like the print function, but in a more elegant fashion.
Browser in R and pdb in Python
In R, there is a function browser() which interrupts the execution and allows the inspection of the current environment. Similarly, there is a module pdb in Python that provides more debugging features. We would only focus on the basic usages of browser() and the set_trace() function in pdb module.
The essential difference between debugging using print() and browser() and set_trace() is that the latter functions allows us to debug in an interactive mode.
Let’s write a function which takes a sorted vector/list v and a target value x as input and returns the leftmost index pos of the sorted vector/list so that v[pos]>=x. Since v is already sorted, we may simply loop through it from left to right to find pos.
chapter2/find_pos.R
1 find_pos=function(v,x){
2 for (i in 1:length(v)){
3 if (v[i]>=x){
4 return(i)
5 }
6 }
7 }
8
9 v=c(1,2,5,10)
10 print(find_pos(v,-1))
11 print(find_pos(v,4))
12 print(find_pos(v,11))
chapter2/find_pos.py
1 def find_pos(v,x):
2 for i in range(len(v)):
3 if v[i]>=x:
4 return i
5
6 v=[1,2,5,10]
7 print(find_pos(v,-1))
8 print(find_pos(v,4))
9 print(find_pos(v,11))
Now let’s run these two scripts.
1 chapter2 $r
2 > source('find_pos.R')
3 [1] 1
4 [1] 3
5 NULL1 chapter2 $python3.7 find_pos.py
2 0
3 2
4 NoneWhen x=11, the function returns NULL in R and None in Python because there is no such element in v larger than x.
The implementation above is trivial, but not efficient. If you have some background in data structures and algorithms, you probably know this question can be solved by binary search. The essential idea of binary search comes from divide-and-conquer5. Since v is already sorted, we may divide it into two partitions by cutting it from the middle, and then we get the left partition and the right partition. v is sorted implies that both the left partition and the right partition are also sorted. If the target value x is larger than the rightmost element in the left partition, we can just discard the left partition and search x within the right partition. Otherwise, we can discard the right partition and search x within the left partition. Once we have determined which partition to search, we may apply the idea recursively so that in each step we reduce the size of v by half. If the length of v is denoted as n, in terms of big O notation6, the run time complexity of binary search is $\mathcal{O}(\log{}n)$, compared with $\mathcal{O}(n)$ of the for-loop implementation.
The code below implements the binary search solution to our question (It is more intuitive to do it with recursion but here I write it with iteration since tail recursion optimization7 in R/Python is not supported).
chapter2/find_binary_search_buggy.R
1 binary_search_buggy=function(v,x){
2 start = 1
3 end = length(v)
4 while (start<end){
5 mid = (start+end) %/% 2 # %/% is the floor division operator
6 if (v[mid]>=x){
7 end = mid
8 }else{
9 start = mid+1
10 }
11 }
12 return(start)
13 }
14 v=c(1,2,5,10)
15 print(binary_search_buggy(v,-1))
16 print(binary_search_buggy(v,5))
17 print(binary_search_buggy(v,11))
chapter2/find_binary_search_buggy.py
1 def binary_search_buggy(v,x):
2 start,end = 0,len(v)-1
3 while start<end:
4 mid = (start+end)//2 # // is the floor division operator
5 if v[mid]>=x:
6 end = mid
7 else:
8 start = mid+1
9 return start
10
11 v=[1,2,5,10]
12 print(binary_search_buggy(v,-1))
13 print(binary_search_buggy(v,5))
14 print(binary_search_buggy(v,11))
Now let’s run these two binary search scripts.
1 chapter2 $r
2 > source( 'binary_search_buggy.R')
3 [1] 1
4 [1] 3
5 [1] 41 chapter2 $python3.7 binary_search_buggy.py
2 0
3 2
4 3The binary search solutions don’t work as expected when x=11. We write two new scripts.
chapter2/find_binary_search_buggy_debug.R
1 binary_search_buggy=function(v,x){
2 browser()
3 start = 1
4 end = length(v)
5 while (start<end){
6 mid = (start+end)
7 if (v[mid]>=x){
8 end = mid
9 }else{
10 start = mid+1
11 }
12 }
13 return(start)
14 }
15 v=c(1,2,5,10)
16 print(binary_search_buggy(v,11))
chapter2/find_binary_search_buggy_debug.py
1 from pdb import set_trace
2 def binary_search_buggy(v,x):
3 set_trace()
4 start,end = 0,len(v)-1
5 while start<end:
6 mid = (start+end)//2
7 if v[mid]>=x:
8 end = mid
9 else:
10 start = mid+1
11 return start
12
13 v=[1,2,5,10]
14 print(binary_search_buggy(v,11))
Let’s try to debug the programs with the help of browser() and set_trace().
1 > source('binary_search_buggy_debug.R')
2 Called from: binary_search_buggy(v, 11)
3 Browse[1]> ls()
4 [1] "v" "x"
5 Browse[1]> n
6 debug at binary_search_buggy_debug.R#3: start = 1
7 Browse[2]> n
8 debug at binary_search_buggy_debug.R#4: end = length(v)
9 Browse[2]> n
10 debug at binary_search_buggy_debug.R#5: while (start < end) {
11 mid = (start + end)%/%2
12 if (v[mid] >= x) {
13 end = mid
14 }
15 else {
16 start = mid + 1
17 }
18 }
19 Browse[2]> n
20 debug at binary_search_buggy_debug.R#6: mid = (start + end)%/%2
21 Browse[2]> n
22 debug at binary_search_buggy_debug.R#7: if (v[mid] >= x) {
23 end = mid
24 } else {
25 start = mid + 1
26 }
27 Browse[2]> n
28 debug at binary_search_buggy_debug.R#10: start = mid + 1
29 Browse[2]> n
30 debug at binary_search_buggy_debug.R#5: (while) start < end
31 Browse[2]> n
32 debug at binary_search_buggy_debug.R#6: mid = (start + end)%/%2
33 Browse[2]> n
34 debug at binary_search_buggy_debug.R#7: if (v[mid] >= x) {
35 end = mid
36 } else {
37 start = mid + 1
38 }
39 Browse[2]> n
40 debug at binary_search_buggy_debug.R#10: start = mid + 1
41 Browse[2]> n
42 debug at binary_search_buggy_debug.R#5: (while) start < end
43 Browse[2]> start
44 [1] 4
45 Browse[2]> n
46 debug at binary_search_buggy_debug.R#13: return(start)
47 Browse[2]> n
48 [1] 4In the R code snippet above, we placed the browser() function on the top of the function binary_search_buggy. Then when we call the function we enter into the debugging environment. By calling ls() we see all variables in the current debugging scope, i.e., v,x. Typing n will evaluate the next statement. After typing n a few times, we finally exit from the while loop because start = 4 such that start < end is FALSE. As a result, the function just returns the value of start, i.e., 4. To exit from the debugging environment, we can type Q; to continue the execution we can type c.
The root cause is that we didn’t deal with the corner case when the target value x is larger than the last/largest element in v correctly.
Let’s debug the Python function using pdb module.
1 chapter2 $python3.7 binary_search_buggy_debug.py
2 > chapter2/binary_search_buggy_debug.py(4) binary_search_buggy()
3 -> start,end = 0,len(v)-1
4 (Pdb) n
5 > chapter2/binary_search_buggy_debug.py(5) binary_search_buggy()
6 -> while start<end:
7 (Pdb) l
8 1 from pdb import set_trace
9 2 def binary_search_buggy(v,x):
10 3 set_trace()
11 4 start,end = 0,len(v)-1
12 5 -> while start<end:
13 6 mid = (start+end)//2
14 7 if v[mid]>=x:
15 8 end = mid
16 9 else:
17 10 start = mid+1
18 11 return start
19 (Pdb) b 7
20 Breakpoint 1 at chapter2/binary_search_buggy_debug.py:7
21 (Pdb) c
22 > chapter2/binary_search_buggy_debug.py(7) binary_search_buggy()
23 -> if v[mid]>=x:
24 (Pdb) c
25 > chapter2/binary_search_buggy_debug.py(7) binary_search_buggy()
26 -> if v[mid]>=x:
27 (Pdb) mid
28 2
29 (Pdb) n
30 > chapter2/binary_search_buggy_debug.py(10) binary_search_buggy()
31 -> start = mid+1
32 (Pdb) n
33 > chapter2/binary_search_buggy_debug.py(5) binary_search_buggy()
34 -> while start<end:
35 (Pdb) start
36 3
37 (Pdb) n
38 > chapter2/binary_search_buggy_debug.py(11) binary_search_buggy()
39 -> return startSimilar to R, command n would evaluate the next statement in pdb. Typing command l would show the current line of current execution. Command b line_number would set the corresponding line as a break point; and c would continue the execution until the next breakpoint (if exists).
In R, besides the browser() function there are a pair of functions debug() and undebug() which are also very handy when we try to debug a function; especially when the function is wrapped in a package. More specifically, the debug function would invoke the debugging environment whenever we call the function to debug. See the example below how we invoke the debugging environment for the sd function (standard deviation calculation).
1 > x=c(1,1,2)
2 > debug(sd)
3 > sd(x)
4 debugging in: sd(x)
5 debug: sqrt(var(if (is.vector(x) || is.factor(x)) x else as.double(x),
6 na.rm = na.rm))
7 Browse[2]> ls()
8 [1] "na.rm" "x"
9 Browse[2]> Q
10 > undebug(sd)
11 > sd(x)
12 [1] 0.5773503The binary_search solutions are fixed below.
chapter2/find_binary_search.R
1 binary_search=function(v,x){
2 if (x>v[length(v)]){return(NULL)}
3 start = 1
4 end = length(v)
5 while (start<end){
6 mid = (start+end)
7 if (v[mid]>=x){
8 end = mid
9 }else{
10 start = mid+1
11 }
12 }
13 return(start)
14 }
chapter2/find_binary_search.py
1 def binary_search(v,x):
2 if x>v[-1]: return
3 start,end = 0,len(v)-1
4 while start<end:
5 mid = (start+end)//2
6 if v[mid]>=x:
7 end = mid
8 else:
9 start = mid+1
10 return start
Benchmarking
By benchmarking, I mean measuring the entire operation time of a piece of program. There is another term called profiling which is related to benchmarking. But profiling is more complex since it commonly aims at understanding the behavior of the program and optimizing the program in terms of time elapsed during the operation.
In R, I like using the microbenchmark package. And in Python, timeit module is a good tool to use when we want to benchmark a small bits of Python code.
As mentioned before, the run time complexity of binary search is better than that of a for-loop search. We can do benchmarking to compare the two algorithms.
chapter2/benchmark.R
1 library(microbenchmark)
2 source('binary_search.R')
3 source('find_pos.R')
4
5 v=1:10000
6
7 # call each function 1000 times;
8 # each time we randomly select an integer as the target value
9
10 # for-loop solution
11 set.seed(2019)
12 print(microbenchmark(find_pos(v,sample(10000,1)), times=1000))
13 # binary-search solution
14 set.seed(2019)
15 print(microbenchmark(binary_search(v, sample(10000,1)), times=1000))
In the R code above, times=1000 means we want to call the function 1000 times in the benchmarking process. The sample() function is used to draw samples from a set of elements. Specifically, we pass the argument 1 to sample() to draw a single element. It’s the first time we use set.seed() function in this book. In R/Python, we draw random numbers based on the pseudorandom number generator (PRNG) algorithm8. The sequence of numbers generated by PRNG is completed determined by an initial value, i.e., the seed. Whenever a program involves the usage of PRNG, it is better to set the seed in order to get replicable results (see the example below).
1 > set.seed(2019)
2 > rnorm(1)
3 [1] 0.7385227
4 > rnorm(1)
5 [1] -0.51476051 > set.seed(2019)
2 > rnorm(1)
3 [1] 0.7385227
4 > set.seed(2019)
5 > rnorm(1)
6 [1] 0.7385227Now let’s run the R script to see the benchmarking result.
1 > source('benchmark.R')
2 Unit: microseconds
3 expr min lq mean median uq max neval
4 find_pos(v, sample(10000, 1)) 3.96 109.5385 207.6627 207.5565 307.8875 536.171 1000
5
6 Unit: microseconds
7 expr min lq mean median uq max neval
8 binary_search(v, sample(10000, 1)) 5.898 6.3325 14.2159 6.6115 7.3635 6435.57 1000The binary_search solution is much more efficient based on the benchmarking result.
Doing the same benchmarking in Python is a bit of complicated.
chapter2/benchmark.py
1 from binary_search import binary_search
2 from find_pos import find_pos
3 import timeit
4 import random
5
6 v=list(range(1,10001))
7
8 def test_for_loop(n):
9 random.seed(2019)
10 for _ in range(n):
11 find_pos(v,random.randint(1,10000))
12
13 def test_bs(n):
14 random.seed(2019)
15 for _ in range(n):
16 binary_search(v,random.randint(1,10000))
17
18 # for-loop solution
19 print(timeit.timeit('test_for_loop(1000)', setup='from __main__ import test_for_loop',number=1))
20 # binary_search solution
21 print(timeit.timeit('test_bs(1000)', setup='from __main__ import test_bs',number=1))
The most interesting part of the Python code above is from __main__ import. Let’s ignore it for now, and we would revisit it later.
Below is the benchmarking result in Python (the unit is second).
1 chapter2 $python3 benchmark.py
2 0.284618441
3 0.00396658900000002Vectorization
In parallel computing, automatic vectorization9 means a program in a scalar implementation is converted to a vector implementation which process multiple pairs of operands simultaneously by compilers that feature auto-vectorization. For example, let’s calculate the element-wise sum of two arrays x and y of the same length in C programming language.
1 int x[4] = {1,2,3,4};
2 int y[4] = {0,1,2,3};
3 int z[4];
4 for (int i=0;i<4;i++){
5 z[i]=x[i]+y[i];
6 }The C code above might be vectorized by the compiler so that the actual number of iterations performed could be less than 4. If 4 pairs of operands are processed at once, there would be only 1 iteration. Automatic vectorization may make the program runs much faster in some languages like C. However, when we talk about vectorization in R/Python, it is different from automatic vectorization. Vectorization in R/Python usually refers to the human effort paid to avoid for-loops. First, let’s see some examples of how for-loops may slow your programs in R/Python.
chapter2/vectorization_1.R
1 library(microbenchmark)
2
3 # generate n standard normal r.v
4 rnorm_loop = function(n){
5 x=rep(0,n)
6 for (i in 1:n) {x[i]=rnorm(1)}
7 }
8
9 rnorm_vec = function(n){
10 x=rnorm(n)
11 }
12
13 n=100
14 # for loop
15 print(microbenchmark(rnorm_loop(n),times=1000))
16 # vectorize
17 print(microbenchmark(rnorm_vec(n),times=1000))
Running the R code results in the following result on my local machine.
1 > source('vectorization_1.R')
2 Unit: microseconds
3 expr min lq mean median uq max neval
4 rnorm_loop(n) 131.622 142.699 248.7603 145.3995 270.212 16355.6 1000
5 Unit: microseconds
6 expr min lq mean median uq max neval
7 rnorm_vec(n) 6.696 7.128 10.87463 7.515 8.291 2422.338 1000 1 import timeit
2 import numpy as np
3
4 def rnorm_for_loop(n):
5 x=[0]*n # create a list with n 0s
6 np.random.seed(2019)
7 for _ in range(n):
8 np.random.normal(0,1,1)
9
10 def rnorm_vec(n):
11 np.random.seed(2019)
12 x = np.random.normal(0,1,n)
13
14 print("for loop")
15 print(f'{timeit.timeit("rnorm_for_loop(100)", setup="from __main__ import rnorm_for_loop",number=1000):.6f} seconds')
16 print("vectorized")
17 print(f'{timeit.timeit("rnorm_vec(100)", setup="from __main__ import rnorm_vec",number=1000):.6f} seconds')Please note that in this Python example we are using the random submodule of numpy module instead of the built-in random module since random module doesn’t provide the vectorized version of random number generation function. Running the Python code results in the following result on my local machine.
Also, the timeit.timeit measures the total time to run the main statements number times, but not the average time per run.
1 chapter2 $python3.7 vectorization_1.py
2 for loop
3 0.258466 seconds
4 vectorized
5 0.008213 secondsIn either R or Python, the vectorized version of random normal random variable (r.v.) is significantly faster than the scalar version. It is worth noting the usage of the print(f'') statement in the Python code, which is different from the way how we print the object of Complex class in \autoref{ch:chp1. In the code above, we use the f-string10 which is a literal string prefixed with 'f' containing expressions inside \{\ which would be replaced with their values. f-string was a feature introduced since Python 3.6. If you are familiar with Scala, you may find that this feature is quite similar with the string interpolation mechanism introduced since Scala 2.10.
It’s also worth noting that lots of built-in functions in R are already vectorized, such as the basic arithmetic operators, comparison operators, ifelse(), element-wise logical operators &,|. But the logical operators &&, || are not vectorized.
In addition to vectorization, there are also some built-in functions which may help to avoid the usages of for-loops. For example, in R we might be able use the apply family of functions to replace for-loops; and in Python the map() function can also be useful. In the Python pandas module, there are also many usages of map/apply methods. But in general the usage of apply/map functions has little or nothing to do with performance improvement. However, appropriate usages of such functions may help with the readability of the program. Compared with the apply family of functions in R, I think the do.call() function is more useful in practice. We would spend some time in do.call() later.
Considering the importance of vectorization in scientific programming, let’s try to get more familiar with vectorization thorough the Biham–Middleton–Levine (BML) traffic model11. The BML model is very important in modern studies of traffic flow since it exhibits a sharp phase transition from free flowing status to a fully jammed status. A simplified BML model could be characterized as follows:
- Initialized on a 2-D lattice, each site of which is either empty or occupied by a colored particle (blue or red);
- Particles are distributed randomly through the initialization according to a uniform distribution; the two colors of particles are equally distributed.
- On even time steps, all blue particles attempt to move one site up and an attempt fails if the site to occupy is not empty;
- On Odd time steps, all red particles attempt to move one site right and an attempt fails if the site to occupy is not empty;
- The lattice is assumed periodic which means when a particle moves out of the lattice, it would move into the lattice from the opposite side.
The BML model specified above is implemented in both R/Python as follows to illustrate the usage of vectorization.
chapter2/BML.R
1 library(R6)
2 BML = R6Class(
3 "BML",
4 public = list(
5 # alpha is the parameter of the uniform distribution to control particle distribution's density
6 # m*n is the dimension of the lattice
7 alpha = NULL,
8 m = NULL,
9 n = NULL,
10 lattice = NULL,
11 initialize = function(alpha, m, n) {
12 self$alpha = alpha
13 self$m = m
14 self$n = n
15 self$initialize_lattice()
16 },
17 initialize_lattice = function() {
18 # 0 -> empty site
19 # 1 -> blue particle
20 # 2 -> red particle
21 u = runif(self$m * self$n)
22 # the usage of L is to make sure the elements in particles are of type integer;
23 # otherwise they would be created as double
24 particles = rep(0L, self$m * self$n)
25 # doing inverse transform sampling
26 particles[(u > self$alpha) &
27 (u <= (self$alpha + 1.0) / 2)] = 1L
28 particles[u > (self$alpha + 1.0) / 2] = 2L
29 self$lattice = array(particles, c(self$m, self$n))
30 },
31 odd_step = function() {
32 blue.index = which(self$lattice == 1L, arr.ind = TRUE)
33 # make a copy of the index
34 blue.up.index = blue.index
35 # blue particles move 1 site up
36 blue.up.index[, 1] = blue.index[, 1] - 1L
37 # periodic boundary condition
38 blue.up.index[blue.up.index[, 1] == 0L, 1] = self$m
39 # find which moves are feasible
40 blue.movable = self$lattice[blue.up.index] == 0L
41 # move blue particles one site up
42 # drop=FALSE prevents the 2D array degenerates to 1D array
43 self$lattice[blue.up.index[blue.movable, , drop = FALSE]] = 1L
44 self$lattice[blue.index[blue.movable, , drop = FALSE]] = 0L
45 },
46 even_step = function() {
47 red.index = which(self$lattice == 2L, arr.ind = TRUE)
48 # make a copy of the index
49 red.right.index = red.index
50 # red particles move 1 site right
51 red.right.index[, 2] = red.index[, 2] + 1L
52 # periodic boundary condition
53 red.right.index[red.right.index[, 2] == (self$n + 1L), 2] = 1
54 # find which moves are feasible
55 red.movable = self$lattice[red.right.index] == 0L
56 # move red particles one site right
57 self$lattice[red.right.index[red.movable, , drop = FALSE]] = 2L
58 self$lattice[red.index[red.movable, , drop = FALSE]] = 0L
59 }
60 )
61 )
Now we can create a simple BML system on a $5\times5$ lattice using the R code above.
1 > source('BML.R')
2 > set.seed(2019)
3 > bml=BML$new(0.4,5,5)
4 > bml$lattice
5 [,1] [,2] [,3] [,4] [,5]
6 [1,] 2 0 2 1 1
7 [2,] 2 2 1 0 1
8 [3,] 0 0 0 2 2
9 [4,] 1 0 0 0 0
10 [5,] 0 1 1 1 0
11 > bml$odd_step()
12 > bml$lattice
13 [,1] [,2] [,3] [,4] [,5]
14 [1,] 2 0 2 1 0
15 [2,] 2 2 1 0 1
16 [3,] 1 0 0 2 2
17 [4,] 0 1 1 1 0
18 [5,] 0 0 0 0 1
19 > bml$even_step()
20 > bml$lattice
21 [,1] [,2] [,3] [,4] [,5]
22 [1,] 0 2 2 1 0
23 [2,] 2 2 1 0 1
24 [3,] 1 0 0 2 2
25 [4,] 0 1 1 1 0
26 [5,] 0 0 0 0 1In the initialization step, we used the inverse transform sampling approach12 to generate the status of each site. Inverse transform sampling method is basic but powerful approach to generate r.v. from any probability distribution given its cumulative distribution function (CDF). Reading the wiki page is enough to master this sampling method.
1 import numpy as np
2
3 class BML:
4 def __init__(self, alpha, m, n):
5 self.alpha = alpha
6 self.shape = (m, n)
7 self.initialize_lattice()
8
9 def initialize_lattice(self):
10 u = np.random.uniform(0.0, 1.0, self.shape)
11 # instead of using default list, we use np.array to create the lattice
12 self.lattice = np.zeros_like(u, dtype=int)
13 # the parentheses below can't be ignored
14 self.lattice[(u > self.alpha) & (u <= (1.0+self.alpha)/2.0)] = 1
15 self.lattice[u > (self.alpha+1.0)/2.0] = 2
16
17 def odd_step(self):
18 # please note that np.where returns a tuple which is immutable
19 blue_index = np.where(self.lattice == 1)
20 blue_index_i = blue_index[0] - 1
21 blue_index_i[blue_index_i < 0] = self.shape[0]-1
22 blue_movable = self.lattice[(blue_index_i, blue_index[1])] == 0
23 self.lattice[(blue_index_i[blue_movable],
24 blue_index[1][blue_movable])] = 1
25 self.lattice[(blue_index[0][blue_movable],
26 blue_index[1][blue_movable])] = 0
27
28 def even_step(self):
29 red_index = np.where(self.lattice == 2)
30 red_index_j = red_index[1] + 1
31 red_index_j[red_index_j == self.shape[1]] = 0
32 red_movable = self.lattice[(red_index[0], red_index_j)] == 0
33 self.lattice[(red_index[0][red_movable],
34 red_index_j[red_movable])] = 2
35 self.lattice[(red_index[0][red_movable],
36 red_index[1][red_movable])] = 0The Python implementation is also given.
1 >>> import numpy as np
2 >>> np.random.seed(2019)
3 >>> from BML import BML
4 >>> bml=BML(0.4,5,5)
5 >>> bml.lattice
6 array([[2, 0, 1, 1, 2],
7 [0, 2, 2, 2, 1],
8 [1, 0, 0, 2, 0],
9 [2, 0, 1, 0, 2],
10 [1, 1, 0, 2, 1]])
11 >>> bml.odd_step()
12 >>> bml.lattice
13 array([[2, 0, 0, 1, 2],
14 [1, 2, 2, 2, 1],
15 [0, 0, 1, 2, 0],
16 [2, 1, 0, 0, 2],
17 [1, 0, 1, 2, 1]])
18 >>> bml.even_step()
19 >>> bml.lattice
20 array([[0, 2, 0, 1, 2],
21 [1, 2, 2, 2, 1],
22 [0, 0, 1, 0, 2],
23 [2, 1, 0, 0, 2],
24 [1, 0, 1, 2, 1]])Please note that although we have imported numpy in BML.py, we import it again in the code above in order to set the random seed. If we change the line to from BML import *, we don’t need to import numpy again. But it is not recommended to import * from a module.
Embarrassingly parallelism in R/Python
According to the explanation of wikipedia13, single-threading is the processing of one command at a time, and its opposite is multithreading. A process is the instance of a computer program executed by one or many threads14. Multithreading is not the same as parallelism. In a single processor environment, the processor can switch execution between threads, which results in concurrent execution. However, it is possible a process with multithreads runs on on a multiprocessor environment and each of the threads on a separate processor, which results in parallel execution.
Both R and Python are single-threaded. In Python, there is a threading package15, which supports multithreading on a single core. It may suit some specific tasks. For example, in web scraping multithreading on a single core may be able to speed up the program if the download time is in excess of the CPU processing time.
Now let’s talk about embarrassingly parallelism by multi-processing. Embarrassingly parallel problem is one where little or no effort is needed to separate the problem into a number of parallel tasks16. In R there are various packages supporting multi-processing on multiple CPU cores, for example, the parallel package, which is my favorite one. In Python, there are also some available modules, such as multiprocessing, joblib and concurrent.futures. Let’s see an application of embarrassingly parallelism to calculate $\pi$ using Monte Carlo simulation17.
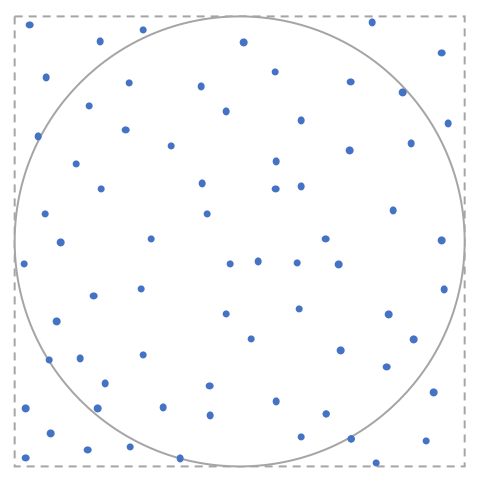
Monte Carlo simulation provides a simple and straightforward way to estimate $\pi$. We know the area of a circle with radius 1 is just $\pi$. Thus, we can convert the original problem of $\pi$ calculation to a new problem, i.e., how to calculate the area of a circle with radius 1. We also know the area of a square with side length 2 is equal to 4. Thus, $\pi$ can be calculated as 4$r_{c/s}$ where $r_{c/s}$ denotes the ratio of areas of a circle with radius 1 and a square with side length 2. Now the problem is how to calculate the ratio $r_{c/s}$? When we randomly throw $n$ points into the square and $m$ of these pionts fall into the inscribed circle, then we can estimate the ratio as $m/n$. As a result, a natural estimate of $\pi$ is $4m/n$. This problem is an embarrassingly parallel problem by its nature. Let’s see how we implement the idea in R/Python.
chapter2/pi.R
1 library(parallel)
2 count_inside_point = function(n){
3 m = 0
4 for (i in 1:n){
5 p_x = runif(1, -1, 1)
6 p_y = runif(1, -1, 1)
7 if (p_x^2 + p_y^2 <=1){
8 m = m+1
9 }
10 }
11 m
12 }
13
14 # now let's use the mcapply for parallelization
15 generate_points_parallel = function(n){
16 # detectCores() returns the number of cores available
17 # we assign the task to each core
18 unlist(mclapply(X = rep(n %/% detectCores(), detectCores()),FUN=count_inside_point))
19 }
20
21 # now let's use vectorization
22 generate_points_vectorized = function(n){
23 p = array(runif(n*2,-1,1), c(n,2))
24 sum((p[,1]^2+p[,2]^2)<=1)
25 }
26
27 pi_naive = function(n) cat('naive: pi -', 4*count_inside_point(n)/n, '\n')
28
29 pi_parallel = function(n) cat('parallel: pi -', 4*sum(generate_points_parallel(n))/n, '\n')
30
31 pi_vectorized = function(n) cat('vectorized: pi -', 4*sum(generate_points_vectorized(n))/n, '\n')
In the above R code snippet, we use the function mclapply which is not currently available on some operation systems18. When it is not available, we may consider to use parLapply instead.
chapter2/pi.py
1 # now let's try the parallel approach
2 # each process uses the same seed, which is not desired
3 def generate_points_parallel(n):
4 pool = mp.Pool()
5 # we ask each process to generate n//mp.cpu_count() points
6 return pool.map(count_inside_point, [n//mp.cpu_count()]*mp.cpu_count())
7
8 # set seed for each process
9 # first, let's define a helper function
10 def helper(args):
11 n, s = args
12 seed(s)
13 return count_inside_point(n)
14
15 def generate_points_parallel_set_seed(n):
16 pool = mp.Pool() # we can also specify the number of processes by Pool(number)
17 # we ask each process to generate n//mp.cpu_count() points
18 return pool.map(helper, list(zip([n//mp.cpu_count()]*mp.cpu_count(), range(mp.cpu_count()))))
19
20 # another optimization via vectorization
21 def generate_points_vectorized(n):
22 p = uniform(-1, 1, size=(n,2))
23 return np.sum(np.linalg.norm(p, axis=1) <= 1)
24
25 def pi_naive(n):
26 print(f'pi: {count_inside_point(n)/n*4:.6f}')
27
28 def pi_parallel(n):
29 print(f'pi: {sum(generate_points_parallel_set_seed(n))/n*4:.6f}')
30
31 def pi_vectorized(n):
32 print(f'pi: {generate_points_vectorized(n)/n*4:.6f}')
In the above Python code snippet, we defined a function generate_points_parallel which returns a list of number of inside points. But the problem of this function is that each process in the pool shares the same random state. As a result, we will obtain the a list of duplicated numbers by calling this function. To overcome the issue, we defined another function generate_points_parallel_set_seed. Let’s see the results of our estimation for $\pi$ running on a laptop.
1 > source('pi.R')
2 > system.time(pi_naive(1e6))
3 naive: pi - 3.144592
4 user system elapsed
5 8.073 1.180 9.333
6 > system.time(pi_parallel(1e6))
7 parallel: pi - 3.1415
8 user system elapsed
9 4.107 0.560 4.833
10 > system.time(pi_vectorized(1e6))
11 vectorized: pi - 3.141224
12 user system elapsed
13 0.180 0.031 0.214 1 >>> import timeit
2 >>> from pi import pi_naive, pi_parallel, pi_vectorized
3 >>> print(f'naive - {timeit.timeit("pi_naive(1000000)", setup="from __main__ import pi_naive",number=1):.6f} seconds')
4 pi: 3.141056
5 naive - 4.363822 seconds
6 >>> print(f'parallel - {timeit.timeit("pi_parallel(1000000)", setup="from __main__ import pi_parallel",number=1):.6f} seconds')
7 pi: 3.141032
8 parallel - 2.204697 seconds
9 >>> print(f'vectorized - {timeit.timeit("pi_vectorized(1000000)", setup="from __main__ import pi_vectorized",number=1):.6f} seconds')
10 pi: 3.139936
11 vectorized - 0.148950 secondsWe see the winner in this example is vectorization, and the parallel solution is better than the naive solution. However, when the problem cannot be vectorized we may use parallelization to achieve better performance.
Evaluation strategy
R is a lazy programming language since it uses lazy evaluation by default19. Lazy evaluation strategy delays the evaluation of an expression until its value is needed. When an expression in R is evaluated, it follows an outermost reduction order in general. But Python is not a lazy programming language and the expression in Python follows an innermost reduction order.
The code snippets below illustrate the innermost reduction vs. outermost reduction.
1 > divide = function(x, y) {stopifnot(y!=0); return(x/y)}
2 > power = function(x, p){if (p==0) 1 else x^p}
3 > power(divide(1,0),2)
4 Error in divide(1, 0) : y != 0 is not TRUE
5 > power(divide(1,0),0)
6 [1] 11 >>> def divide(x, y): return x/y
2 >>> def power(x, p): return 1 if p==0 else x**p
3 >>> power(divide(1,0),0)
4 Traceback (most recent call last):
5 File "<stdin>", line 1, in <module>
6 File "<stdin>", line 1, in divide
7 ZeroDivisionError: division by zeroBecause of the outermost reduction order, the R code snippet evaluate the function power first and since if second argument is 0 the first argument is not required to evaluate. Thus, the function call returns 1. But the Python code snippet first evaluates the function call of divide and an exception is raised because of division by zero.
Although Python is an eager language, it is possible to simulate the lazy evaluation behavior. For example, the code inside a generator20 is evaluated when the generator is consumed but not evaluated when it is created. We can create a sequence of Fibonacci numbers with the help of generator.
1 >>> def fib():
2 ... '''
3 ... a generator to create Fibonacci numbers less than 10
4 ... '''
5 ... a, b = 0, 1
6 ... while a<10:
7 ... a, b = b, a+b
8 ... yield a
9 ...
10 >>> f = fib()
11 >>> print(f)
12 <generator object fib at 0x116dfc570>
13 >>> for e in f: print(e)
14 ...
15 1
16 1
17 2
18 3
19 5
20 8
21 13
22 >>> In the code snippet above, we create a generator which generates the sequence of Fibonacci numbers less than 10. When the generator is created, the sequence is not generated immediately. When we consume the generator in a loop, each element is then generated as needed. It is also common to consume the generator with the next function.
1 >>> f = fib()
2 >>> next(f)
3 1
4 >>> next(f)
5 1
6 >>> next(f)
7 2
8 >>> next(f)
9 3
10 >>> next(f)
11 5Speed up with C/C++ in R/Python
The purpose of this section is not to teach C/C++. If you know how to write C/C++ then it may help to improve the performance of R/Python program.
It is recommended to use vectorization technique to speed up your program whenever possible. But what if it’s infeasible to vectorize the algorithm? One potential option is to use C/C++ in the R/Python code. According to my limited experience, it is much easier to use C/C++ in R with the help of Rcpp package than in Python. In Python, more options are available but not as straightforward as the solution of Rcpp. In this section, let’s use Cython21. Actually, Cython itself is a programming language written in Python and C. In Rcpp we can write C++ code directly and use it in native R code. With the help of Cython, we can write python-like code which is able to be compiled to C or C++ and automatically wrapped into python-importable modules. Cython could also wrap independent C or C++ code into python-importable modules.
Let’s see how to calculate Fibonacci numbers with these tools. The first step is to install Rcpp package in R and the Cython module (use pip) in Python. Once they are installed, we can write the code in C++ directly for R. As for Cython, we write the python-like code which would be compiled to C/C++ later.
chapter2/Fibonacci.cpp
1 #include <Rcpp.h>
2 using namespace Rcpp;
3
4 // [[Rcpp::export]]
5 int Fibonacci(int n) {
6 if (n==1 && n==2){
7 return 1;
8 }
9 return Fibonacci(n-1)+Fibonacci(n-2);
10 }
chapter2/Fibonacci.pyx
1 #!python
2 #cython: language_level=3
3
4 cdef int Fibonacci_c(int n):
5 if n==1 or n==2:
6 return 1
7 return Fibonacci_c(n-1) + Fibonacci_c(n-2)
8
9 def Fibonacci_static(n):
10 return Fibonacci_c(n)
11
12 def Fibonacci(n):
13 if n==1 or n==2:
14 return 1
15 return Fibonacci(n-1) + Fibonacci(n-2)
It’s worth noting the extension of the Cython file above is pyx, not py. And in Fibonacci.pyx, the function we defined is quite following the native Python syntax. But in Cython we can add static typing declarations which are often helpful to improve the performance, although not mandatory. The keyword cdef makes the function Fibonacci_c invisible to Python and thus we have to define the Fibonacci_static function as a wrapper which can be imported into Python. The function Fibonacci is not static typed for benchmarking.
Let’s also implement the same functions in native R/Python for benchmarking purpose.
chapter2/Fibonacci.R
1 Fibonacci_native = function(n){
2 if (n==1 || n==2){
3 1
4 }else {
5 Fibonacci_native(n-1) + Fibonacci_native(n-2)
6 }
7 }
chapter2/Fibonacci.py
1 def Fibonacci_native(n):
2 if n==1 or n==2:
3 return 1
4 return Fibonacci_native(n-1) + Fibonacci_native(n-2)
Now let’s compare the performance of different implementations.
1 > library(microbenchmark)
2 > library(Rcpp)
3 > source("Fibonacci_native.R")
4 > sourceCpp('Fibonacci.cpp')
5 > # the native implementation in R
6 > microbenchmark(Fibonacci_native(20), times=1000)
7 Unit: milliseconds
8 expr min lq mean median uq max neval
9 Fibonacci_native(20) 3.917138 4.056468 4.456824 4.190078 4.462815 26.30846 1000
10 > the C++ implementation
11 > microbenchmark(Fibonacci(20), times=1000)
12 Unit: microseconds
13 expr min lq mean median uq max neval
14 Fibonacci(20) 14.251 14.482 15.81708 14.582 14.731 775.095 1000The native implementation in R is much slower the C++ implementation (see the difference of units).
1 >>> import pyximport
2 >>> pyximport.install()
3 (None, <pyximport.pyximport.PyxImporter object at 0x1056f9f98>)
4 >>> from Fibonacci import Fibonacci_static, Fibonacci
5 >>> from Fibonacci_native import Fibonacci_native
6 >>> import timeit
7 >>> print(f'{timeit.timeit("Fibonacci_native(20)", setup="from __main__ import Fibonacci_native",number=1000):.6f} seconds')
8 1.927359 seconds
9 >>> print(f'{timeit.timeit("Fibonacci(20)", setup="from __main__ import Fibonacci",number=1000):.6f} seconds')
10 0.316726 seconds
11 >>> print(f'{timeit.timeit("Fibonacci_static(20)", setup="from __main__ import Fibonacci_static",number=1000):.6f} seconds')
12 0.012741 secondsThe results show the static typed implementation is the fastest, and the native implementation in pure Python is the slowest. Again the time measured by timeit.timeit is the total time to run the main statement 1000 times. The average time per run of Fibonacci_static function is close to the average time per run of Fibonacci in R.
A first impression of functional programming
Functional (programming) languages define programs as mathematical functions and treat them as first-class2]. In pure functional programming, a function takes input and returns output without making any side effect. A side effect means a state variable outside the function is modified. For example, printing the value of a variable inside a function is a side effect.
Both R and Python are not purely functional languages per se, although R behaves more functional than Python23. If you wonder why the return function could be ignored to return a value in an R function, now you get the explanation – a function should return the output automatically from a FP perspective.
I don’t think it makes any sense to debate on if we should choose OOP or FP using R/Python. And both languages support multiple programming paradigms (FP, OOP etc.). If you want to do purist FP, neither R nor Python would be the best choice. For example, purist FP should avoid loop because loop always involves an iteration counter whose value changes over iterations, which is obviously a side effect. Thus, recursion is very important in FP. But tail recursion optimization is not supported in R and Python although it can be implemented via trampoline24.
Let’s introduce a few tools that could help to get a first impression of FP in R and Python.
Anonymous function
We have seen anonymous functions before. Anonymous functions are commonly used in FP. To recap, let’s see how to define a useless anonymous function.
1 > function(x) x*x
2 function(x) x*x1 >>> lambda x: x**2
2 <function <lambda> at 0x1033d5510>Usually, we define anonymous functions because we want to pass them to other functions. And thus, the functions defined above are useless as they are not passed to any other function and after the definition they are deleted automatically from the memory.
Of course, we can assign names to the anonymous functions – but what is the point to name anonymous functions?
1 > f1 = function(x) x*x
2 > f1
3 function(x) x*x1 >>> f1 = lambda x: x**2
2 >>> f1(2)
3 4Map
Map could be used to avoid loops. The first argument to map is a function.
In R, we use Map which is shipped with base R; and in Python we use map which is also a built-in function.
1 > Map(function(x) x*x, c(1:3))
2 [[1]]
3 [1] 1
4
5 [[2]]
6 [1] 4
7
8 [[3]]
9 [1] 9
10 > Map(function(x, y) x*y, c(1:3), c(4:6))
11 [[1]]
12 [1] 4
13
14 [[2]]
15 [1] 10
16
17 [[3]]
18 [1] 181 >>> map(lambda e: e**2, [1,2,3])
2 <map object at 0x1033e3160>
3 >>> list(map(lambda x: x**2, [1,2,3]))
4 [1, 4, 9]
5 >>> list(map(lambda x, y: x*y, [1,2,3], [4,5,6]))
6 [4, 10, 18]There are a few things to note from the code snippets above. First, the returned value from Map in R is a list rather than a vector. That is because Map is just a wrapper of mapply function in R, and the SIMPLIFY argument is set to FALSE. If we want to get a vector value returned, just use mapply instead.
1 > mapply(function(x, y) x*y, c(1:3), c(4:6), SIMPLIFY=TRUE)
2 [1] 4 10 18The returned value from map in Python is a map object which is an iterator. The generator function we have seen before is also an iterator. To use an iterator, we could convert it to a list, or get the next value using the next function, or just use a for loop.
1 >>> # use next on an iterator
2 >>> o=map(lambda x, y: x*y, [1,2,3], [4,5,6])
3 >>> next(o)
4 4
5 >>> next(o)
6 10
7 >>> o=map(lambda x, y: x*y, [1,2,3], [4,5,6])
8 >>> # use for loop on an iterator
9 >>> for e in o:print(e)
10 ...
11 4
12 10
13 18
14 >>> # convert an iterator to a list
15 >>> o=map(lambda x, y: x*y, [1,2,3], [4,5,6])
16 >>> list(o)
17 [4, 10, 18]Filter
Similar to map, the first argument to Filter in R and filter in Python is also a function. This function is used to get the elements which satisfy a certain condition specified in the first argument. For example, we can use it to get the even numbers.
1 > Filter(function(x) x%%2==0, c(0:10))
2 [1] 0 2 4 6 8 101 >>> list(filter(lambda e: e%2==0, range(11)))
2 [0, 2, 4, 6, 8, 10]Reduce
Reduce behaves a bit different from Map and Filter. The first argument is a function with exactly two arguments. The second argument is a sequence, on each element of which the first function argument will be applied from left to right. And there is one optional argument called initializer. When the initializer is provided, it would be used as the first argument of the function argument of Reduce. The examples below depict how it works.
1 > Reduce(function(x, y) x+2*y, c(1,2,3))
2 [1] 11
3 > Reduce(function(x, y) x+2*y, c(1,2,3), 1)
4 [1] 131 >>> from functools import reduce
2 >>> reduce(lambda x, y:x+2*y, [1,2,3])
3 11
4 >>> reduce(lambda x, y:x+2*y, range(1,4), 1)
5 13Please note that in order to use reduce in Python, we have to import it from the functools module.
Utilizing these functions flexibly may help to make the code more concise, at the cost of readability.
Miscellaneous
We have introduced the basics of R/Python programming so far. There are much more to learn to become an advanced user of R/Python. For example, the appropriate usages of iterator, generator, decorator could improve both the conciseness and readability of your Python code. The generator25 is commonly seen in machine learning programs to prepare training/testing samples. decorator is a kind of syntactic sugar to allow the modification of a function’s behavior in a simple way. In R there are no built-in iterator, generator, decorator, but you may find some third-party libraries to mimic these features; or you may try to implement your own.
One advantage of Python over R is that there are some built-in modules containing high-performance data structures or commonly-used algorithms implemented efficiently. For example, I enjoy using the deque structure in the Python collections module26, but there is no built-in counterpart in R. We have written our own binary search algorithm earlier in this Chapter, which can also be replaced by the functions in the built-in module bisect27 in Python.
Another important aspect of programming is testing. Unit testing is a typical software testing method that is commonly adopted in practice. In R there are two third-party packages testthat and RUnit. In Python, the built-in unittest is quite powerful. Also, the third-party module pytest28 is very popular.
1 https://www.rstudio.com/products/rstudio
2 https://code.visualstudio.com/
3 https://www.jetbrains.com/pycharm
4 https://docs.python.org/3/library/logging.html
5 https://en.wikipedia.org/wiki/Divide-and-conquer_algorithm
6 https://en.wikipedia.org/wiki/Big_O_notation
7 https://en.wikipedia.org/wiki/Tail_call
8 https://en.wikipedia.org/wiki/Pseudorandom_number_generator
9 https://en.wikipedia.org/wiki/Automatic_vectorization
10 https://www.python.org/dev/peps/pep-0498
11 https://en.wikipedia.org/wiki/Biham-Middleton-Levine_traffic_model
12 https://en.wikipedia.org/wiki/Inverse_transform_sampling
13 https://en.wikipedia.org/wiki/Thread_(computing)
14 https://en.wikipedia.org/wiki/Process_(computing)
15 https://docs.python.org/3.7/library/threading.html
16 https://en.wikipedia.org/wiki/Embarrassingly_parallel
17 https://en.wikipedia.org/wiki/Monte_Carlo_method
18 https://stat.ethz.ch/R-manual/R-devel/library/parallel/doc/parallel.pdf
19 Floréal Morandat, Brandon Hill, Leo Osvald, and Jan Vitek. Evaluating the design of the r language. In European Conference on Object-Oriented Programming, pages 104–131. Springer, 2012.
20 https://www.python.org/dev/peps/pep-0255
21 https://en.wikipedia.org/wiki/Cython
22 https://en.wikipedia.org/wiki/List_of_programming_languages_by_type#Functional_languages
23 John M Chambers et al. Object-oriented programming, functional programming and r. Statistical Science, 29(2):167–180, 2014.
24 https://en.wikipedia.org/wiki/Tail_call#Through_trampolining
25 https://docs.python.org/3/howto/functional.html
26 https://docs.python.org/3/library/collections.html
27 https://docs.python.org/3.7/library/bisect.html
28 https://docs.pytest.org/en/latest